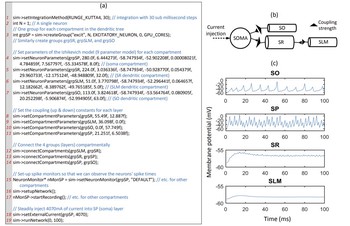
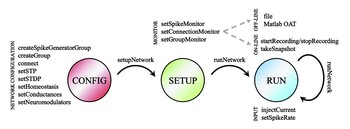
We have developed CARLsim 4, a user-friendly SNN library written in C++ that can simulate large biologically detailed neural networks. Improving on the efficiency and scalability of earlier releases, the present release allows for the simulation using multiple GPUs and multiple CPU cores concurrently in a heterogeneous computing cluster. …
Topic: CARLsim
Research Projects
CARLsim 4: An open source library for large scale, biologically detailed spiking neural network simulation using heterogeneous clusters
Ting-Shou Chou, Hirak J. Kashyap, Jinwei Xing, Stanislav Listopad, Emily L. Rounds, Michael Beyeler, Nikil Dutt, Jeffrey L. Krichmar Proceedings of the 2018 International Joint Conference on Neural Networks (IJCNN)
CARLsim 3: A user-friendly and highly optimized library for the creation of neurobiologically detailed spiking neural networks
We have developed CARLsim 3, a user-friendly, GPU-accelerated SNN library written in C/C++ that is capable of simulating biologically detailed neural models. The present release of CARLsim provides a number of improvements over our prior SNN library to allow the user to easily analyze simulation data, explore synaptic plasticity rules, and automate …
Michael Beyeler, Kristofor D. Carlson, Ting-Shou Chou, Nikil Dutt, Jeffrey L. Krichmar Proceedings of the 2015 International Joint Conference on Neural Networks (IJCNN)